M.R.AI
Alzheimer's detection from MRI scans — a team comparison of four CNN configurations on the OASIS dataset, with patient-level splits to avoid leakage.
Why
A five-person team project for the Biomedical Image Processing with AI course at UFV (December 2025). The professor offered several disease-detection problems; we picked Alzheimer’s because the trade-offs were the most interesting — a small, imbalanced dataset, and the open question of how much pretrained CNN backbones can actually do on medical imaging without specialised pipelines.
What we built
A controlled comparison of four configurations on the OASIS Alzheimer’s MRI dataset (Kaggle), all evaluating the same binary task (Demented vs. Non-Demented):
| Full scan | Center-cropped (truncated) | |
|---|---|---|
| ResNet18 (PyTorch) | AUC 0.911 ✱ | AUC 0.859 |
| VGG16 (Keras) | AUC 0.651 | AUC 0.812 |
✱ Best configuration.
How it works
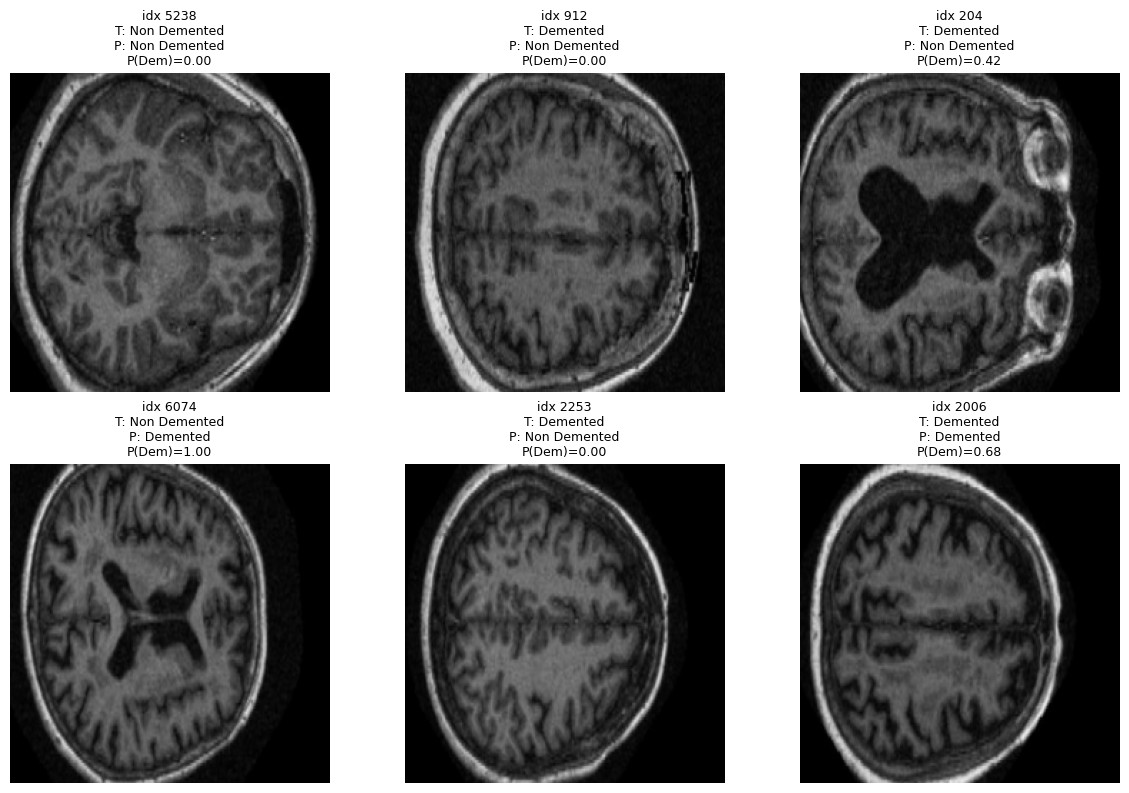
Data. OASIS Alzheimer’s Detection (Kaggle), built on OASIS-1: ~80,000 T1 structural MRI slices from 461 patients, originally labelled in four classes (Non Demented, Very Mild, Mild, Moderate Dementia). We collapsed those into a binary task — Demented vs. Non Demented — to fit a clear clinical decision and reduce ambiguity between adjacent stages.
Two safeguards before training: we balanced at the patient level, capping Non Demented to 120 patients (from 266) to avoid the dataset’s natural skew toward healthy controls; and we split at the patient level, not the image level — same patient never appears across train/val/test. This avoids the common leakage trap where the model memorises patients instead of pathology.
Preprocessing. Two variants compared in parallel: full scan resized to 224×224, vs. center-cropped to 160×160 (keeping the central brain region and discarding skull/edges) then resized to 224. Grayscale → 3-channel, ImageNet normalisation. Standard augmentation on training: rotation, horizontal flip, resized crop.
Models. Two pretrained backbones, fine-tuned:
- ResNet18 (PyTorch, ImageNet weights). Frozen except
layer4and a custom classification head (Dropout + Linear → 2 classes). - VGG16 (Keras, ImageNet weights). Frozen except the top, with GlobalAveragePooling + Dense head.
Training. Adam + scheduler, early stopping on validation loss. Same training budget for all four configurations to keep the comparison fair.
Results
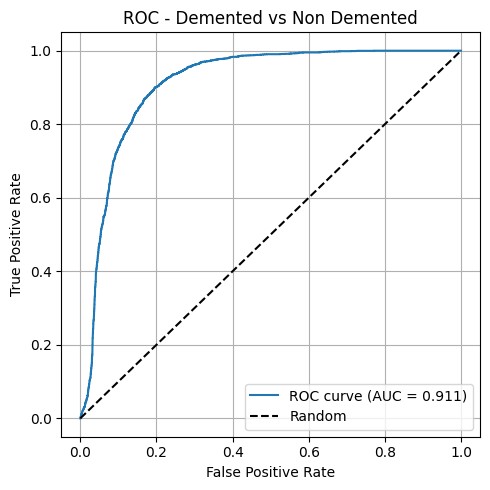
The cleanest finding wasn’t the absolute numbers — it was that the effect of cropping depends on the architecture. Truncating the input damaged ResNet18 (0.911 → 0.859) but rescued VGG16 (0.651 → 0.812). The likely reason is that ResNet18 uses global context efficiently and the cropped scan removes signal; VGG16 struggles with the full scan and benefits from a tighter region of interest.
Best single model: ResNet18 on full scans, AUC = 0.911, 78.2% accuracy on the held-out patient-split test set (F1 Demented = 0.67, F1 Non-Demented = 0.72, Mean IoU = 0.611).
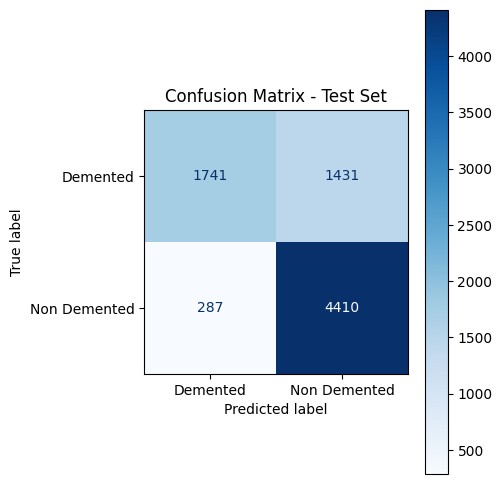
The confusion matrix is honest about the asymmetry: the model is conservative — it under-calls dementia (1,431 false negatives vs. 287 false positives). For a screening tool that’s the wrong direction; for a teaching exercise it’s a useful failure mode to discuss.
What’s not here
We wanted to add Grad-CAM saliency maps to check whether the model attends to clinically meaningful regions (hippocampus, ventricles) or spurious correlations. We didn’t get to it inside the course deadline. Same for proper class-balanced loss and LIME/SHAP comparisons. Listed here so it’s clear what the project is and isn’t.
Status
Prototype. Academic team project (5 people) at UFV, December 2025. Pitched as part of the CloseAI startup-pitch academic project.
See it
- Live site: mrai-gray.vercel.app
- Code: github.com/RAvila-bioeng/M.R.AI (team repo, MIT license)
- Contact: hello@damianvidalc.com